 WE researchers often joke that no-one ever publishes negative results, but that doesn’t mean to say that negative results aren’t extremely useful. On one level, knowledge of such negative results can prevent you repeating the same mistakes that countless other researchers, in other labs, have undoubtedly made over the years. On the other hand, they can provide a valuable dataset with which to generate new and useful information. One such example is the ‘Negatome Database‘, which has been reported by Smialowski et al.1 in Nucleic Acids Research advance access (November 17, 2009).
WE researchers often joke that no-one ever publishes negative results, but that doesn’t mean to say that negative results aren’t extremely useful. On one level, knowledge of such negative results can prevent you repeating the same mistakes that countless other researchers, in other labs, have undoubtedly made over the years. On the other hand, they can provide a valuable dataset with which to generate new and useful information. One such example is the ‘Negatome Database‘, which has been reported by Smialowski et al.1 in Nucleic Acids Research advance access (November 17, 2009).
The Negatome is a collection of protein and domain (functional units of proteins) pairs that are unlikely to be engaged in direct physical interactions. But why on Earth would we want to know about proteins that don’t interact with each other; in fact, why do we need to know about proteins that interact at all?
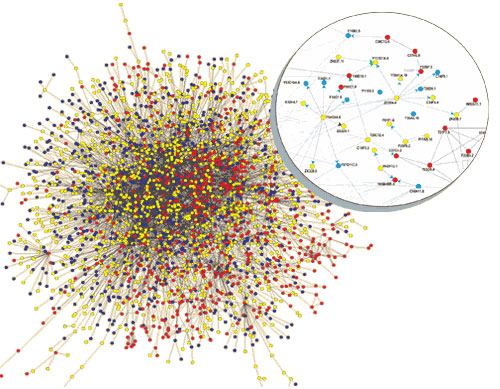
 Researchers recognize that that a cell doesn’t function purely by the action of individual proteins, but instead by large macromolecular complexes mediated by many interacting proteins. The image to the left indicates an example macromolecular ‘machine’, in this case those involved in signal processing at the neuronal synapses (and which are likely to be working quite hard right now!).
Researchers recognize that that a cell doesn’t function purely by the action of individual proteins, but instead by large macromolecular complexes mediated by many interacting proteins. The image to the left indicates an example macromolecular ‘machine’, in this case those involved in signal processing at the neuronal synapses (and which are likely to be working quite hard right now!).
Understanding protein-protein interactions is critical to understanding the biochemistry that makes us tick, and promises invaluable information about how such interactions change in disease processes, or when a bacterial cell becomes resistant to antibiotics, or a human cancer cell resistant to a particular chemotherapy treatment. These studies, called interactomics, are a branch of the relatively new field of systems biology, and add a new layer of information to the raw data collected as a result of projects such as the human genome project.
“Understanding the human genome definitely does not go far enough to explain what makes us different from more simple creatures,” says Professor Michael Stumpf at Imperial. “Our study indicates that protein interactions could hold one of the keys to unravelling how one organism is differentiated from another.” – Author of ‘Estimating the size of the human interactome’ 2 [Wellcome news].
Current C. elegans (worm) interactome, the most complete interactome so far (via The Scientist).
Obviously there is a need for such studies, but why do we need a negatome?
There are disparities in the reported estimated number of interacting proteins in the human interactome, with some estimating 130,000 binary protein interactions 3, and others indicating 650,000 binary protein interactions 2. These may well reflect the difference between those proteins that can interact (i.e. biophysical interactions), and those that do interact (i.e. biological interactions).
The big problems with studies investigating interactomics is the noise. Noise is unwanted and erroneous data that obfuscates the true interactions that researchers hope to observe. Sometimes, when looking for an interaction between a particular protein with one of thousands of other proteins, you sometimes get false discoveries where the proteins appear to interact even though they shouldn’t; similairly, you also get false negatives, and you don’t want to miss out on an interaction that is happening.
In fact, a study 4 has attempted to quantitate just how many false discoveries and false negatives actually affect interactomic studies:
False discoveries: Yeast (9.9 %), worm (13.2 %) and fly (17 %).
False negatives: Yeast (51 %), worm (42 %) and fly (28 %).
Controls are therefore essential in such studies, to rule out the noise. An extensive ‘gold-standard’ dataset of positive interactions exists for some model organisms; these are pooled from careful literature curation and provide the basis upon which computer algorithms are trained to recognise the characteristics of a statistically probable interaction.
“The lack of negative training data represents a significant problem because the knowledge about NIPs [non-interating proteins] is as important for developing and evaluating prediction algorithms as the knowledge of true positive pairs.“
Hence the facility of the negatome database. Smialowski et al. constructed the negatome database using two approaches:
1. The collection of evidence against physical interactions from literature, focusing only on those cases where the lack of interaction between two proteins was experimentally validated by an individual experiment. – Thus empirical evidence for there being no interaction.
2. Through analysis of complexes consisting of three or more proteins deposited in the PDB (Protein Data Bank), derived a set of protein pairs that, while being in immediate vicinity in the context of a protein complex, do not interact directly with each other. – The protein databank is a store of known protein crystal structures that are produced by mixing the component proteins at reasonably high concentrations, in a specially determined chemical environment, in order for them to form crystals of protein. If two proteins don’t interact under such conditions, as seen from the final crystal structure determination, then there’s a good chance that they really don’t interact.
The database currently provides a total of 1892 non-interacting proteins and 979 predicted non-interacting domain (function units of proteins) pairs based on the experimental evidence. As such, the negatome is well on the way to become a ‘gold-standard’ dataset for training predictors of protein–protein interaction, thus proving that negative results are still results.
—-
1 Smialowski, P., Pagel, P., Wong, P., Brauner, B., Dunger, I., Fobo, G., Frishman, G., Montrone, C., Rattei, T., Frishman, D., & Ruepp, A. (2009). The Negatome database: a reference set of non-interacting protein pairs Nucleic Acids Research DOI: 10.1093/nar/gkp1026
2 Stumpf, M., Thorne, T., de Silva, E., Stewart, R., An, H., Lappe, M., & Wiuf, C. (2008). From the Cover: Estimating the size of the human interactome Proceedings of the National Academy of Sciences, 105 (19), 6959-6964 DOI: 10.1073/pnas.0708078105
3 Venkatesan, K., Rual, J., Vazquez, A., Stelzl, U., Lemmens, I., Hirozane-Kishikawa, T., Hao, T., Zenkner, M., Xin, X., Goh, K., Yildirim, M., Simonis, N., Heinzmann, K., Gebreab, F., Sahalie, J., Cevik, S., Simon, C., de Smet, A., Dann, E., Smolyar, A., Vinayagam, A., Yu, H., Szeto, D., Borick, H., Dricot, A., Klitgord, N., Murray, R., Lin, C., Lalowski, M., Timm, J., Rau, K., Boone, C., Braun, P., Cusick, M., Roth, F., Hill, D., Tavernier, J., Wanker, E., Barabási, A., & Vidal, M. (2008). An empirical framework for binary interactome mapping Nature Methods, 6 (1), 83-90 DOI: 10.1038/nmeth.1280
4 Huang, H., & Bader, J. (2008). Precision and recall estimates for two-hybrid screens Bioinformatics, 25 (3), 372-378 DOI: 10.1093/bioinformatics/btn640
Expect a visit from the Omics Police soon! Good luck! ^_^
Cheers,
-Psi-
Don’t even think about it! I’m just working with what pithy jargon keeps getting published lol
I hate Omics as much as any researcher who’s worked in molecular genetics for a decade, but playing lip service to this interrupted the flow.
…thanks for stopping by 😉
Interesting I wrote a letter about the plague of ‘omics’ in 2004, so here come the cops
http://www.the-scientist.com/2005/02/14/8/4/
More seriously, the whole field seems to me a bit of a mess. How many of these interactions actually occur in living cells? Even if they were all real, what can you do with them if you don’t know their association and disocciation rate constants for their interaction, and their concentrations? Well you can make a powerpoint with multicoloured blobs, join with lines that have no numbers. Then what?
You can say “protein interactions could hold one of the keys to unravelling how one organism is differentiated from another”, but that must be truism of the decade.
Let’s hope that one day it becomes science.
Thanks for your comment David.
My own stand point, having worked in high through-put approaches to determining interactions in Staphylococcus aureus, is that it is a process of whittling down to something of substance. Initial huge, high stringency screens would be performed to isolate particular interacting groups, which would flag those for study by slightly more labour intensive approaches, followed by further refinement and, as you say, some kinetic measurements. With it being a real chore discovering that many of the interactions are false hits, it’d be nice, in theory, to narrow these down somewhat. Alas, the utility of such databases to the world of S. aureus is likely dubious; it’s not one of the hot topics, though I feel it should be. But then don’t we all about our pet organisms?
In many of the protein-DNA and protein-protein interactions on which I’ve worked it’s true to say that in vitro determinations of on and off rates has provided little useful information because in vivo there has been compartmentalisation, and localised concentrations of proteins mediated by their interaction at DNA binding sites or transmembrane Type IV secretory systems. Thus there is of course a limitation to any biochemical investigation.
I have a love/hate relationship with systems biology. I consider it partly to blame for their huge grants hogging the lion’s share of recent funding opportunities in which, in any other year, I might have gotten funded. But on the other hand, it’s about time some of the biochemistry was put back into context with the overall network of biochemical interactions. If databases formulated from empirical data derived from direct determinations of such interactions (or via the PBD) can be of use in achieving this goal, then I’m all for them trying.
p.s. I’ve also said elsewhere, not wanting to be defensive of Omics in any way (ahem, and not just because I’ve blogged about it), but I think the ‘Negatome’, being a database, is a piece of surprisingly good punnery from a German group, being a ‘tome’ of negative interaction information for ‘interactomics’.
Or maybe I’m affording them too much wit?
Kill the ‘interactomics’ if needs be 😉
RE the Omics Police: They came.
Congrats on your new award! (well, not really yours per se, but you brought the word to light so… 😉 )
I agree that ‘interactomics’ was way worse, but it’s also a rather bland abuse, unworthy of any prize… negatome is just…kind of awesome!
lol thanks for that! (I think) 😉